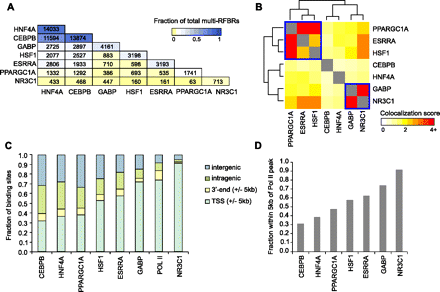
Genome-wide patterns of binding and colocalization of PPARGC1A and its network partners. (A) Pairwise colocalization of transcriptional regulators within multi-RFBRs. Numbers in each cell represent the number of multi-RFBRs that are co-occupied by the corresponding pair of regulators. The first cell in each column indicates the total number of multi-RFBRs bound by each regulator. Cells are colored by the fraction of all 16,712 multi-RFBRs identified in the genome. Factors are ordered by decreasing number of multi-RFBRs bound. (B) Heatmap depicting the clustering of transcriptional regulators based on the similarity of their colocalization patterns. Colors in the heatmap indicate the strength of association between each pair of regulators (odds ratio from a Fisher's exact test; see Methods). Two groups of regulators exhibiting highly similar colocalization patterns based on the results of clustering are outlined in blue boxes. (C) Distribution of transcriptional regulator binding sites relative to UCSC Known Genes transcript annotations (Hsu et al. 2006). Factors are ordered by increasing proportion of TSS-adjacent binding sites. (D) Bar graph depicting the fraction of binding sites within 5 kb of an RNA pol II peak for each regulator.











