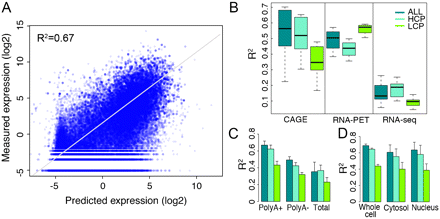
Accuracy of the TF model for predicting TSS expression levels. (A) Consistency of predicted values with expression levels measured by CAGE in Poly A+ RNA samples extracted from whole cells. (B) Comparison of predictive accuracies of the TF model for expression data generated by three different technologies: CAGE, RNA–PET, and RNA-seq. (C) Comparison of predictive accuracies of the TF model for expression data from three different RNA extraction protocols: Poly A+, Poly A-, and total RNA. (D) Comparison of predictive accuracies of the TF model for expression data in different cellular components. In B–D, only data sets from K562 are used. The binding signals of 40 TFSSs are used as predictors. HCP and LCP are high and low CpG content promoters, respectively. Separate models are constructed for ALL, HCP, and LCP categories.











