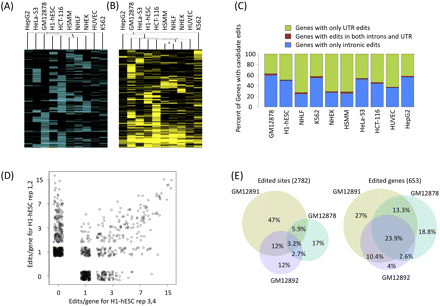
Gene level analysis of RNA editing after private SNV filtering. (A) Hierarchical clustering of the editing frequency of the 33.5% (1905 out of 5695 possible) individual A-to-G candidate editing sites occurring in at least two distinct cell types. (B) Hierarchical clustering of the number of edits in the 47.4% (662 out of 1395 possible) of genes edited in at least two distinct cell types. (C) RNA editing in genes cluster in the UTR or in the introns with few genes having edits in both UTR and introns. Percentage of genes with only UTR edits are in green; intronic edits, blue; and edits in both introns and UTR, red. (D) Reproducibility of calling RNA edits for human H1 ES cells. Scatter plot of RNA edit calls for rep 1,2 versus rep 3,4 is on a log2-log2 scale with a pseudocount of 1. A Gaussian noise was added to points to visualize density. (E) Venn diagrams of A-to-G candidate edits in lymphoblastoid cells from a hapmap trio. The Venn diagram of the individual sites (left) and edited genes (right); 35.8% of the union of edited sites are found in two or more cell types, while 54.2% of the union of edited genes are found in two or more cell types.











