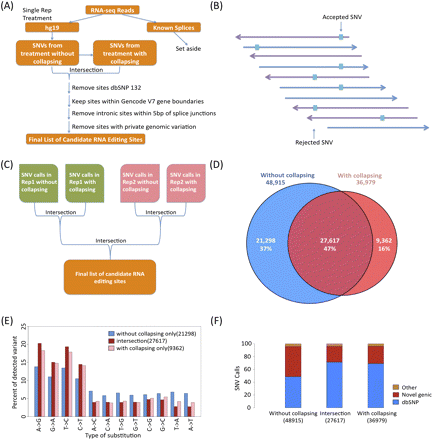
RNA SNV calling strategy. (A) Flowchart of analysis: 75-bp paired-end RNA-seq reads were mapped onto an extended genome (genome + known splice junctions + spikes) using Bowtie. Reads mapping onto splice sites and spikes were set aside, and reads mapping onto hg19 were used to call single nucleotide variants (SNVs). A parallel set of analyses was done using a collapsed set of reads with unique coordinates, and the intersections of SNVs from the uncollapsed and collapsed treatments were obtained. Known SNPs annotated in dbSNP132, sites outside gene boundaries, and intronic sites within 5 bp of splice junctions were removed. For the GM trio, any candidate with evidence of a private genomic variation was also removed. (B) Example of candidate editing site. Purple arrows pointing to the left represent reads on the (−) strand, while blue arrows pointing to the right represent reads on the (+) strand. The blocks represent variants between the reference DNA and the RNA-seq. A SNV is kept when at least three nonidentical reads support the SNV, with a minimum SNV frequency of 10%, and at least one edit per strand. (C) Intersection strategy for two replicates. For cell types with two replicates, the SNVs remaining after collapsing were intersected between the replicates. (D) The number of SNVs remaining after collapsing for the prefiltered sites. Number of SNVs that are only in the uncollapsed set are in blue; the intersection, purple; and collapsed set, red. (E) Collapsing increases the relative amount of A-to-G SNVs and also increases the relative number of transitions. Number of SNVs that are only in the uncollapsed set are in blue; the intersection, purple; and collapsed set, red. (F) The fraction of dbSNP is highest in the intersection of the full and collapsed sets. The relative amount of calls found in dbSNP132, novel genic SNVs, and other SNVs in the uncollapsed set are at the left; the collapsed set, right; and the intersection of the two, middle.











