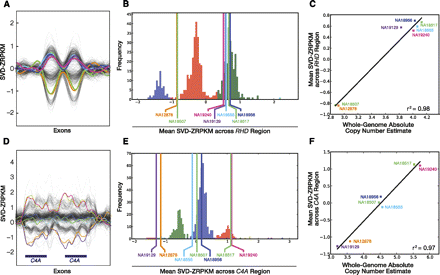
CNP locus genotyping of RHD and C4A. (A) SVD-transformed values for exons for the Rhesus deletion factor locus (RHD/RHCE) show distinct copy number states across both paralogous genes. (B) Histogram of average SVD-ZRPKM values for the ESP data set (533 individuals) and seven HapMap samples. Clustering was performed using an unsupervised algorithm (Supplemental Note). (C) Correlation between SVD-ZRPKM genotype values (y-axis) and absolute copy number estimate (x-axis) based on whole-genome read-depth for seven HapMap samples and experimentally validated by array-CGH. (D–F) Similar to above, for C4A locus.











