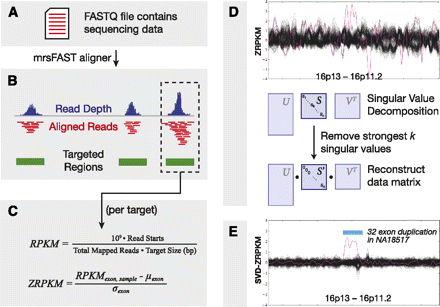
Method overview and CNV discovery. Exome sequencing reads from FASTQ files were divided into nonoverlapping 36-bp constituents (A) and aligned to targeted regions (B), allowing for up to two mismatches per 36-bp alignment. (C) For each exon or targeted region, we calculated RPKM values and then transformed these into “ZRPKM” values based on the median and standard deviation of each exon across all samples. (D) ZRPKM values were inputted into the SVD transformation, where we removed the first 12–15 singular values. Finally, a centrally weighted 15-exon average was passed over the SVD-ZRPKM values in order to reduce false positives, and a ±1.5 SVD-ZRPKM threshold was used to discover CNVs. (E) Final image shows ZRPKM values from 1000 consecutive exons on chromosome 16, plotted for 533 ESP exome background samples (black traces) and NA18507 (pink trace). Blue bar corresponds to a rare duplication in NA18507 at the METTL9/OTOA locus at chr16p12.2 that was validated by SNP microarray CNV analysis.











