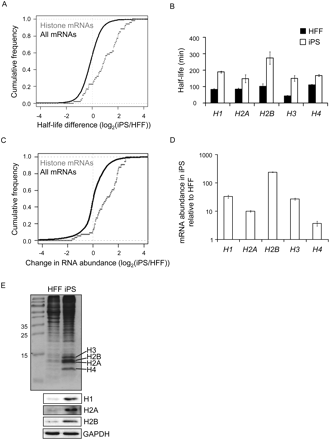
Histone expression is increased in iPS cells. (A) Graph comparing the change in half-life for 53 histone mRNAs in HFF and iPS cells to changes for all 5481 mRNAs for which half-lives were calculated (P-value 2.2 × 10−11). (B) Graph showing the half-life of individual histone mRNAs in HFF and iPS cells as determined by qRT-PCR. Error bars denote standard deviation derived from three replicates. (C) Graph comparing the change in abundance of 53 histone mRNAs to that of all 19,190 mRNAs expressed in both iPS and HFF cells. These data were derived from the 0-min time point of the half-life experiment (P-value 6.5 × 10−9). (D) Graph showing increases in abundance of individual histone mRNAs as determined by qRT-PCR. Error bars denote the standard deviation derived from three replicates. (E, top) Coomassie-stained gel showing abundance of histone proteins in extracts derived from equal numbers of HFF and iPS cells. (Bottom) Western blots showing abundance of Histones H1, H2A, and H2B in extracts derived from equal numbers of HFF and iPS cells.











