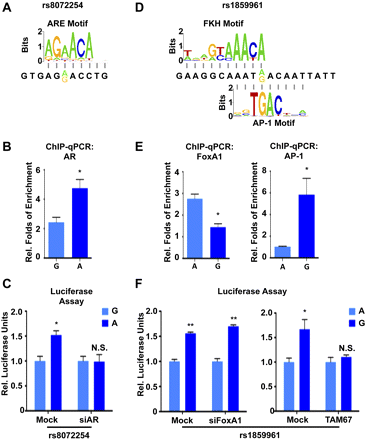
The functional SNPs modulate transcription factor bindings, further imposing allele-specific gene expression. (A) The PWM matrixes of ARE DNA motif overlapping with rs8072254 are indicated. The alleles of rs8072254 are presented; (yellow) reference G allele; (green) variant A allele. (B) In vitro allele-specific AR ChIP-qPCR assay of the two alleles from rs8072254. DNA enrichment is normalized to the DNA input. (Y-axis) Relative folds of enrichment with regard to FOXA1 ± 1 SEM. The P-value is derived from a t-test; (*) p ≤ 0.05. (C) A luciferase assay was performed in PCa LNCaP cells. The experiments were transfected with control (Mock) and AR siRNA, respectively. Luciferase units were normalized to Renilla signal and further normalized to the G allele of rs8072254 ± 1 SEM. The P-value is derived from a t-test; (*) p ≤ 0.05; (N.S.) non-significant. (D) The PWM matrixes of FKH and AP-1 DNA motif overlapping with rs1859961 are indicated. The alleles of rs1859961 are presented; (green) reference A allele; (yellow) variant G allele. (E) Same as B but with regard to FOXA1 and AP-1 ChIP-qPCR assay of the two alleles from rs1859961; (*) p ≤ 0.05. (F) A luciferase assay was performed in PCa LNCaP cells. (Left panel) The experiments were transfected with Control (Mock) and FOXA1 siRNA, respectively. (Right panel) The experiments were treated with an empty control plasmid and the dominant-negative AP-1 mutant TAM67 plasmid, respectively. Luciferase units were normalized to Renilla signal, and further normalized to the A allele of rs1859961 ± 1 SEM. The P-value is derived from a t-test; (*) p ≤ 0.05; (**) p ≤ 0.01; (N.S.) non-significant.











