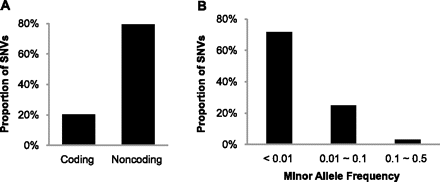
Observed patterns of population polymorphisms that are consistent with the neutral theory of molecular evolution. (A) Proportions of single nucleotide variants (SNVs) with a rejected substitution (RS) score greater than 3 in coding and noncoding regions (Goode et al. 2010). A lower degree of genetic variation is permitted by purifying selection in functionally important regions (coding) of the genome as compared with noncoding regions with no known function. (B) Proportions of nonsynonymous SNVs predicted to be damaging by SIFT and PolyPhen-2 programs (Marth et al. 2011), which show an enrichment of deleterious (nonsynonymous) variants among the rare polymorphisms (allele frequency <1%).











