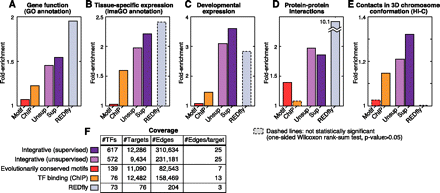
Functional enrichment and coverage of regulatory networks. Functional enrichment of coregulated genes for annotation with similar functions (A), colocalization in tissues (B), coexpression in an independent developmental time-course (C), physical interactions between coregulated genes (D), and colocalization in three-dimensional chromosome conformation (E). (F) Coverage of the networks. The manually curated REDfly network shows strong enrichment for coregulated genes to be annotated with similar functions, colocalized in tissues, coexpressed in development and to physically interact, thus supporting the relevance of these metrics, although it did not pass the significance test in B,C, and E due to its small size. The integrative networks show significant enrichment for all five metrics, while including ∼10-fold more TFs and ∼100-fold more targets than REDfly. In contrast, the two physical networks (conserved motifs and ChIP binding) show relatively lower enrichment.











