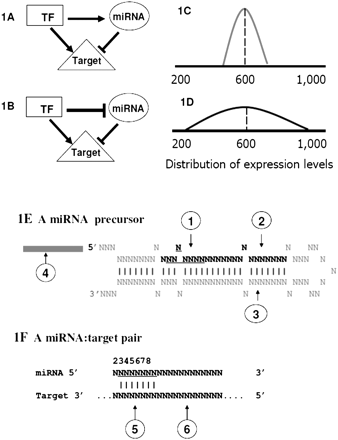
Scheme for a canonical miRNA regulation network. The comprehensive interactions between miRNAs and transcription factors (TF) are expected to comprise “wired” genetic networks to regulate the expression of target genes. (A,B) Examples of incoherent and coherent feed-forward loops, respectively. In the incoherent feed-forward loop, the direct regulatory effect of a TF on the target gene is opposed to the indirect regulatory effect through miRNA regulation (A); and in the coherent feed-forward loops, the direct regulatory effect of a TF on the target gene is synergistic to the indirect regulatory effect through miRNA regulation (B). The consequence of miRNA regulatory effects is to reduce the stochastic noise in expression levels of target genes. (C,D) Distributions of expression levels (the x-axis is the number of molecules per cell) of two hypothetical target genes across cells (or individuals). (C) Expression levels of the target gene are tightly regulated due to miRNA targeting; (D) expression levels of a gene not targeted by miRNAs are highly variable across individuals. (E,F) Six sources of genetic variation in the miRNA regulatory networks. (E) The scheme of a miRNA precursor characterized by a hairpin structure. The sequence in black is the mature miRNA (guide strand) and position 2–8 of mature miRNA is the “seed” region (underlined). Perfect pairing between the seed of the mature miRNA and the target site is crucial for miRNA target recognition (F). Mutations associated with a miRNA precursor can be divided into four categories: (1) mutations in the “seed” alter the target recognition; (2) changes in the mature miRNA beyond the “seed” region potentially affect target recognition; (3) changes outside mature miRNAs can affect miRNA biogenesis and hence affect the abundance of the mature miRNA; and (4) changes in promoter regions of the miRNA precursor will cause the abundance of mature miRNA to be variable (E). Mutations in miRNA target pairing regions also affect miRNA binding: (5) Mutations in the seed pairing region of a miRNA will affect the target recognition and hence the expression level of the host genes; and (6) mutations in regions of 3′ UTR beyond seed pairing might affect the accessibility of a miRNA to the target site. In our model, genetic variation in the former four classes is defined as the “trans-regulatory” effect, and variation in the latter two classes is defined as the “cis-regulatory” effect. Both trans- and cis-regulatory effects in the miRNA regulatory networks contribute to the expression variation of the target genes.











