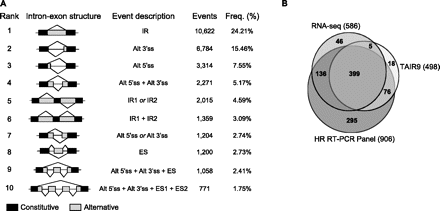
Alternative splicing events and validation of assembled transcripts. (A) Top 10 most frequent types of AS in the predicted transcripts according to ASTALAVISTA. The first column illustrates the intron-exon structure of the AS event, followed by its description, the raw number of events found in the sample, and their frequency. (IR) Intron retention, (Alt 5′ss) alternative 5′ splice site, (Alt 3′ss) alternative 3′ splice site, (ES) exon skipping. (B) Venn diagram of the number of fragments obtained in the HR RT-PCR panel, putative assembled transcripts of RNA-seq, and TAIR9-annotated transcripts according to primer pairs of the HR RT-PCR panel (see text and Methods).











