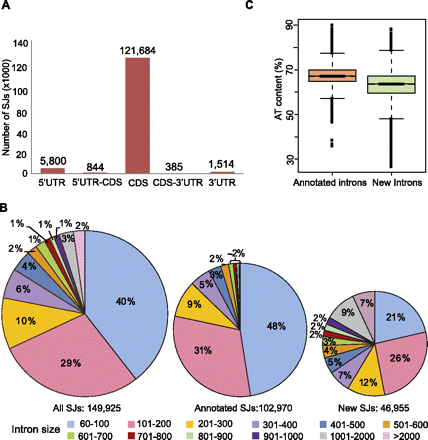
Features of splice junctions and predicted introns. (A) Distribution of splice junctions along gene features in protein-coding genes, according to TAIR9. (B) Distribution of intron sizes defined by splice junctions (SJ) predicted by TopHat (left circle). Splice junctions were classified as annotated if they were already in TAIR9 (middle circle); otherwise, they were classified as new (right circle). (C) Distribution of the AT content (%) in introns generated by annotated splice junctions and new splice junctions.











