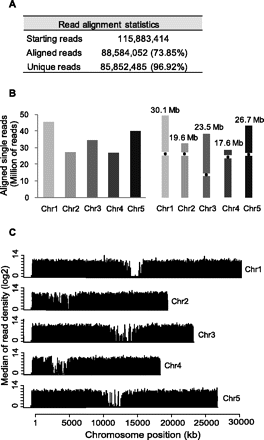
Figure 1.
Read alignment using Bowtie and TopHat. (A) Table showing statistics of the aligned reads to the Arabidopsis genome (TAIR9). (B) Aligned single reads to Arabidopsis chromosomes (left) and schematic representation of Arabidopsis chromosomes (right). (C) Log2 scale of median read density in windows of 1 kb by chromosome.











