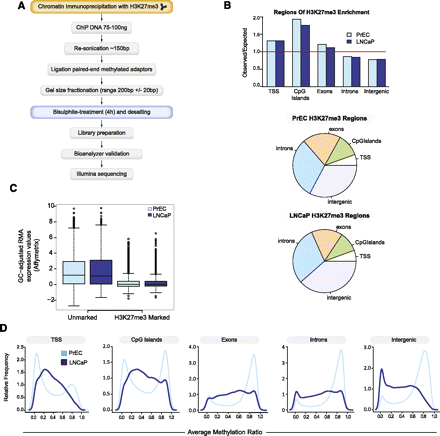
BisChIP-seq DNA methylation profiles of H3K27me3-enriched DNA from normal PrEC and cancer LNCaP cells. (A) Flowchart of BisChIP-seq protocol to perform bisulfite treatment and library preparation on H3K27me3-ChIP DNA. (B) Distribution of H3K27me3-enrichment genome-wide relative to observed over expected and pie charts showing relative distributions across the genome. (C) Affymetrix Gene 1.0 ST expression values for H3K27me3-marked and -unmarked genes in PrEC and LNCaP cells. (D) Distribution frequency of CpG methylation levels at H3K27me3-marked regions that fall into each regional annotation category from low (0%) to high (100%) methylation (0.0–1.0).











