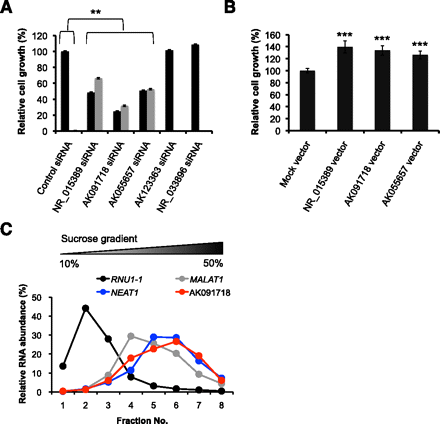
Functional analysis of SLiTs. (A) HeLa TO cells were treated with a control siRNA or with siRNAs targeting indicated SLiTs. The number of viable cells was counted using the Cell Counting Kit-8. Two siRNAs were used for NR_015389, AK091718, and AK055657 to rule out the possibility of an off-target effect of siRNA (black or gray bars indicate the first or second siRNA, respectively). The relative abundance (100% in control cells) is shown in the graph. Values represent mean ± SD obtained in four replicate experiments [(**) P < 0.01, Student's t-test]. (B) HeLa TO cells were treated with plasmid vectors as indicated. The number of viable cells was counted using the Cell Counting Kit-8. The relative abundance (100% with mock vector) is shown in the graph. Values represent mean ± SD in six replicate experiments [(***) P < 0.001, Student's t-test]. (C) The relative amount of RNA was quantified by RT-qPCR from nuclear fractions obtained by sucrose gradient centrifugation.











