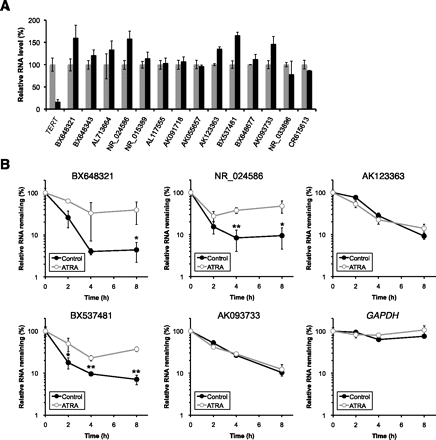
Degradation of several SLiTs was altered by all-trans retinoic acid (ATRA) in HeLa TO cells. (A) Altered abundance of SLiTs by ATRA except BC018860, whose level was below the detection limit. HeLa TO cells were untreated (gray bar) or treated with 10 mM ATRA (black bar). Values represent mean ± errors obtained from duplicate experiments. (B) Decay rates of 5 SLiTs, whose levels were increased >135% in ATRA-treated cells compared with control cells in A and of GAPDH were determined by BRIC and RT-qPCR in control cells (solid circle and black bar) and in ATRA-treated cells (open circle and gray bar). The relative quantitative values at time 0 h were arbitrarily adjusted to 100%. Values represent mean ± SD obtained from triplicate experiments [(**) P < 0.01; (*) P <0.05, Student's t-test].











