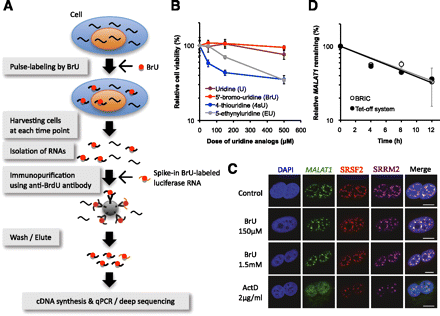
5′-Bromo-uridine immunoprecipitation chase (BRIC) for measuring RNA decay in physiologically undisturbed conditions in vivo. (A) Overview of BRIC. (B) A549 cells were treated with various uridine analogs at the concentrations indicated for 48 h. The number of viable cells was counted using the Cell Counting Kit-8 (Wako). The relative abundance (100% in untreated cells) is shown in the graph. Values represent mean ± SD obtained from four replicate experiments. (C) A549 cells were treated with 5′-bromo-uridine (BrU) or Actinomycin D (ActD) at the concentrations indicated for 7 h. RNA-FISH was used to detect MALAT1 (green). Nuclear speckle proteins, SRSF2 (red) and SRRM2 (magenta), were assayed by immunostaining. Cells were counterstained with DAPI (scale bars, 10 μm). (D) Decay rates of MALAT1 were determined by BRIC and RT-qPCR (gray line) or by the Tet-off system and RT-qPCR (black line) in HeLa TO cells. Relative quantitative values at time 0 h were arbitrarily adjusted to 100%. Values represent mean ± errors obtained from duplicate experiments. The half-lives of MALAT1 using BRIC or the Tet-off system were 7.6 h and 7.0 h, respectively.











