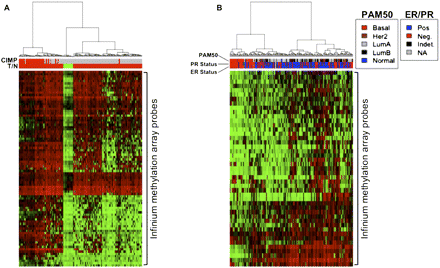
Hierarchical cluster analysis of primary TCGA colon and breast cancer samples based on Infinium β-values of PRC-module genes identified as methylated in colon (A) or breast (B) cancer cell lines. (A) Colon tumors are classified as CIMP+ (top red bars in the heat map) if eight or more of a 12-CIMP marker panel (Weisenberger et al. 2006) are methylated in a tumor sample, and otherwise classified as CIMP− (top gray bars in the heat map). (Red) Tumor and (green) normal (T/N) samples. (B) Breast tumor samples were classified as estrogen (ER)/progesterone (PR) positive or negative and also classified according to their gene expression (PAM50) classes. The PRC-module genes sufficiently discern the CIMP+ and CIMP− tumors in both colon and breast samples and tightly cluster important breast cancers subtypes that are independently defined based on gene expression signatures.











