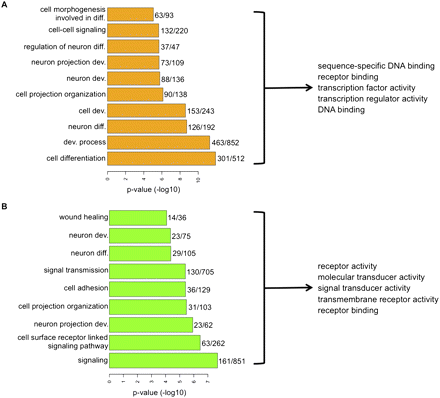
Cancer-specific hypermethylated genes are enriched for functions associated with developmental regulation and comprise a list of genes that are tightly silenced in tumor cells compared with MSCs. Enrichment of biological processes in hypermethylated genes marked by bivalency (A) or H3K4Me3 (B) in ESCs against a background list of genes that, respectively, have the same mark and are present on the methylation array. (Right) Top five molecular function categories of the genes constituting the significantly enriched biological processes. The numbers adjacent to the bars are the total number of genes in the bivalent or H3K4Me3-marked genes in that category (denominator) and the number of genes in the category that are DNA hypermethylated in 50% or more of all the cancers examined (numerator). Supplemental Tables 2 and 3 give the gene lists used for GO analysis and all the top categories identified.











