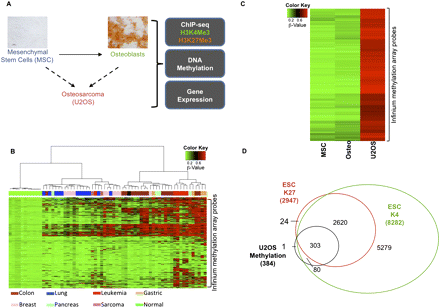
Genes with promoter-proximal CpG hypermethylation in osteosarcoma are greatly enriched for a bivalent chromatin history in ESCs and are down-regulated in osteosarcoma cells compared with ESCs. (A) Schematic of the experimental design. von Kossa staining shows differentiation of MSCs to osteoblasts (scale bar, 50 μm). (B) Heat map of β-values for 2489 Infinium probes (Supplemental Table 1) within CpG islands of 1891 genes for cell lines corresponding to different tumor types. These are probes that are methylated in at least one cell line and not methylated in any of the normal cells (see Methods). Different cell types are shown below the plot. (C) Heat map of the β-values for MSCs, osteoblasts, and U2OS cells. Genes with β-value >0.75 in U2OS and <0.25 in MSCs and osteoblasts were selected as hypermethylated genes in osteosarcoma. (D) Extent of overlap of the osteosarcoma-hypermethylated genes with genes marked by H3K4Me3 or H3K27Me3 in ESCs. Osteosarcoma-hypermethylated genes overlap significantly with ESC-bivalent genes (P-value < 0.001).











