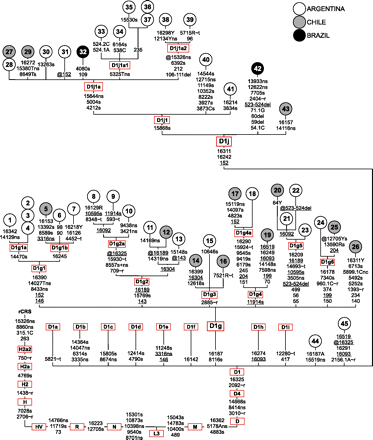
Phylogenetic tree of southern South American haplogroups D1g and D1j. This tree includes 43 new complete mtDNA sequences belonging to D1g and D1j and illustrates subhaplogroup affiliations. The position of the revised Cambridge Reference Sequence (rCRS) (Andrews et al. 1999) is indicated for reading off sequence motifs. Samples #1–#26 belong to D1g, while samples #27–#43 belong to D1j. The country of origin of the sample donors is indicated by circle color (see legend). All SNPs and indels are shown on the branches except for cytosine insertions at np 309. In the case of transversions, insertions, or heteroplasmic mutations, the base is indicated according to the IUPAC nucleotide code. The prefix @ indicates the reversion of a mutation occurring earlier in the phylogeny. The suffixes “s” and “ns” indicate synonymous and nonsynonymous substitutions, respectively, while “∼t” and “∼r” indicate affected positions in tRNA and rRNA loci, respectively. Recurrent mutations within the phylogeny are underlined. The mutational motifs of the previously described D1 subhaplogroups (D1a–D1f, D1h, D1i) are also shown (van Oven and Kayser 2009) (Build 13). D1 sequences #44 and #45 are also new. Additional information regarding each of the 45 novel mitochondrial genomes is available in Supplemental Tables S1 and S2.











