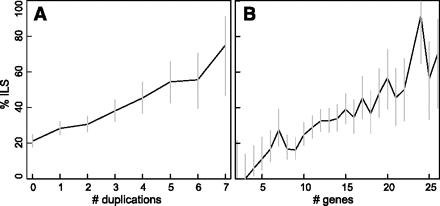
Duplications increase the rate of incomplete lineage sorting (ILS). Using the DLCoal model, we simulated 2000 gene trees for the 12 flies phylogeny, using an effective population size of N = 5 × 106, duplication-loss rates of λ = μ = 0.0048 events/gene per million years, and 10 generations/yr. (A) As more gene duplications occur in a gene tree, the probability of ILS increases. (B) Overall, larger gene families tend to have increased ILS frequency. Error bars indicate 95% confidence intervals.











