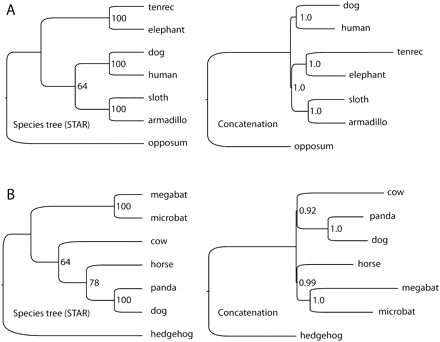
Species trees and concatenated trees from high genomic coverage data sets for two rapid radiations in placental mammals. (A) Species tree from a 591-locus analysis identifies Afrotheria as the first-diverging lineage of placental mammals, whereas alternate topologies had less than half the bootstrap support (Table 1). A Bayesian analysis based on concatenated data places Afrotheria and Xenarthra together with high PP. Data on gene-tree discordance from Figure 2 suggest that this may be because the basal divergence of placental mammals lies close to the phylogenetic anomaly zone. (B) Species tree of taxa in the Laurasiatheria based on 683 loci places bats (Chiroptera) in a traditional location sister to Perissodactyla, Cetartiodactyla, and Carnivora. Bayesian analysis based on concatenated data produced an unusual tree with bats grouping sister to Perissodactyla, but with Carnivora grouping with Cetartiodactyla. The species tree and concatenated analysis of the 183 locus data set produced a different topology, more supportive of the hypothesized clade Pegasoferae (see text), suggesting that a robust understanding of this divergence event will require further investigation incorporating additional taxa and loci. Note that STAR trees do not contain branch length information.











