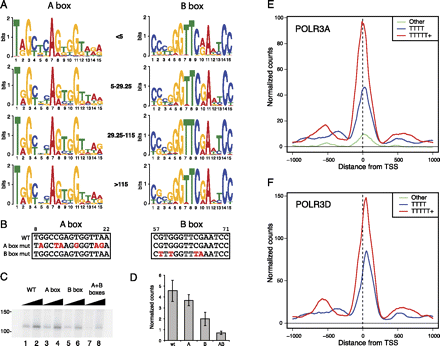
The qualities of the A and B box and terminator sequences correlate with RNAP-III occupancy. (A) The LOGOs obtained for the A and B boxes in tRNA genes with the RNAP-III scores indicated in the middle. (B) A and B box mutations introduced into the serine tRNA gene (chr11_1821_tRNASer_GCT-). Note that bases 3–6, 14, and 15 of the A box, and bases 2–6, 14, and 15 of the B box are, for most tRNAs, involved in base-pairing in the clover leaf structure. For the tRNA gene analyzed here, the bases involved in base-pairing are bases 3–5 of the A box and bases 2–6, 14, and 15 of the B box. (C) In vitro transcription with the serine tRNA gene, either wild type, or with mutations in the A, B, or both A and B boxes as indicated above the lanes. (D) Quantitation of six replicate experiments as in C. (E) Average tag density profiles for POLR3A in genes with strong terminators (5Ts or more, red line), medium terminators (4Ts, blue line), or poor terminators (less than 4Ts, green line). (F) As in E, but for POLR3D.











