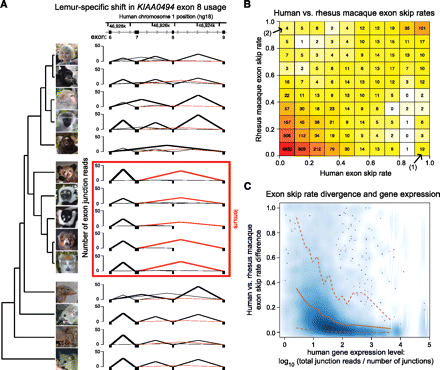
Exon structure divergence and evolution. (A) Phylogenetic shift in splicing and exon usage in the KIAA0494 gene. For each species, the y-axis depicts the number of RNA-seq reads spanning junctions of exons 6–9 (x-axis) based on human reference genome exon positions. Lines representing the number of reads spanning the exon 7 to 9 junction, observed in the overwhelming majority of inferred transcripts in lemurs but only rarely in other species, are highlighted in red. Junctions representing the most common transcript in each species are bolded. (B) Extreme divergence in exon skipping is rare. We mapped our RNA-seq read data against the human and rhesus macaque reference genome sequences to assess patterns of exon usage divergence independently of our assembled gene database (see Supplemental Methods). Shown is a heatmap depicting human vs. rhesus macaque exon skip rates. Included in this plot are all exons with at least 10 reads covering junctions, summed across all individuals of both species, and at least eight reads entering, exiting, or skipping the exon in each species. The number of exons with significant, complete divergence skip rates (i.e., exons always skipped in one species and never skipped in the other; three total), are shown by arrows in the upper left and lower right boxes of the heatmap. (C) Density plot comparing the absolute difference in human versus rhesus macaque exon skip rates to estimated expression levels (human) for the gene containing that exon, for all identified exons with evidence of alternative splicing or differential exon usage, regardless of expression level. Mean and 95th/fifth percentiles are depicted as solid and dashed red lines, respectively. Lower-expressed genes are more likely to harbor exons with larger between-species exon usage differences, reflecting either statistical artifacts or relatively lower constraint on exon structure and splicing on lower-expressed genes, or both.











