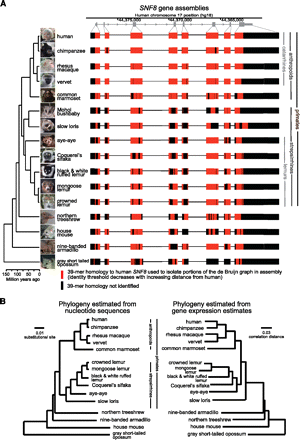
Transcript assembly and phylogenetic reconstruction from RNA-seq data. (A) Typical example of an assembled gene, SNF8, with complete cross-species exon conservation. (Red bars) Identified homologies to the human SNF8 RefSeq coding sequence that were used to isolate the appropriate region of the de Bruijn graph during the assembly process. Divergence times are approximate and based on consensus estimates from previous studies. Photos of strepsirrhine primates were kindly provided by David Haring, Duke Lemur Center. (B) Neighbor-joining trees estimated from nucleotide sequence and gene expression data. Nucleotide sequence distance matrix was computed from concatenated multispecies alignments of coding sequences of 515 genes that were assembled for all 16 species. Gene expression pairwise correlation distance matrix was computed for species mean expression estimates using all genes assembled in at least six species (6494 genes). As expected, the known primate phylogeny was recapitulated perfectly from the nucleotide sequence data (see Supplemental Fig. S7 for the tree, also including bushbaby), with the only discrepancy among nonprimate mammals being the juxtaposition of the mouse and armadillo branches, likely explained by long branch attraction that is a common issue in phylogenetic analyses that include rodents (Cannarozzi et al. 2007). Variation in the expression data also follows a phylogenetic pattern but with slow loris erroneously placed outside all other primates and the misplacement of armadillo.











