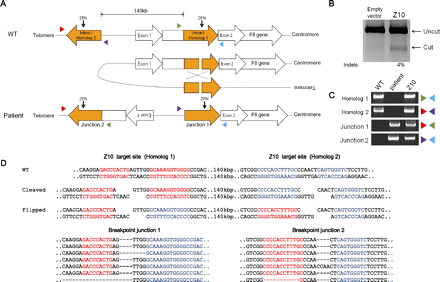
Targeted inversion of the F8 gene using ZFNs. (A) Schematic representation of a chromosomal inversion that causes severe hemophilia A. NAHR between two homologous regions, one in intron 1 of the F8 gene (here named homolog 1) and the other located 140 kbp upstream (homolog 2), gives rise to an inversion found in severe hemophilia A. The two homologous regions are oriented in opposite directions. PCR primers (colored triangles) used to detect the inversion are shown. Z10 target sites are indicated by arrows. (B) Site-specific mutations at the Z10 target site in the F8 intron 1 revealed by the T7 endonuclease assay. (C) PCR products corresponding to wild-type and inversion genotypes. Genomic DNA isolated from a hemophilia A patient was used as a positive control for the inversion-specific PCR. (D) DNA sequences of breakpoint junctions of the inversion events. Z10 target sites are shown in red (homolog 1) or blue (homolog 2).











