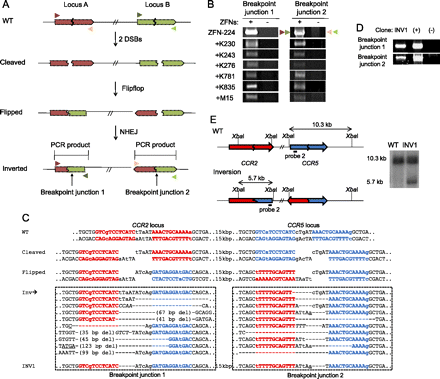
Inversions induced by ZFNs. (A) Schematic representation of ZFN-mediated inversions. Zigzag lines indicate ZFN target sites. PCR primers (triangles) used for the detection of two breakpoint junctions that result from genomic inversions are shown. (B) PCR products corresponding to inversion events in cells treated with various combinations of two ZFN pairs. (C) DNA sequences of breakpoint junctions of the inversion events. Nucleotide sequences of CCR5 and CCR2 sites are shown in blue and red, respectively. (Inv) Various inversion junctions induced by ZFN-224. Inversion junction sequences induced by other ZFNs are shown in Supplemental Figure 4. The two breakpoint junction sequences in the INV1 clone are also shown. Nonconserved bases at the CCR2 and CCR5 loci are shown in lowercase letters. Symbols are as in Figure 1. (D) PCR products validating inversions in the INV1 clone. (+) and (−) are as in Figure 2D. (E) Schematic of CCR2 and CCR5 loci in the wild-type and inversion alleles and Southern blot analysis. Symbols are as in Figure 1.











