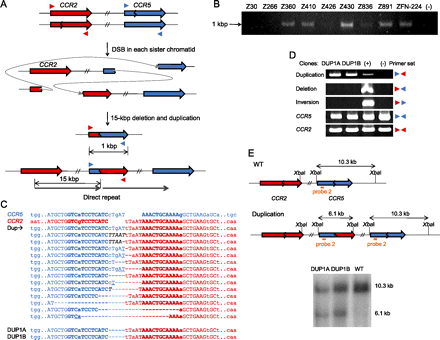
Duplications induced by ZFNs. (A) Schematic representation of ZFN-mediated duplications. Zigzag lines indicate ZFN target sites. Colored triangles indicate approximate positions of PCR primers. (B) PCR products corresponding to the 15-kbp duplications in cells treated with various ZFNs. (−) indicates a negative control (cells transfected with empty plasmid). (C) DNA sequences of breakpoint junctions of the duplications. Nucleotide sequences of CCR5 and CCR2 sites are shown in blue and red colors, respectively. Dup indicates various duplication junctions induced by ZFN-224. Duplication junction sequences induced by other ZFNs are shown in Supplemental Figure 3. DNA sequences of breakpoint junctions in two clones, DUP1A and DUP1B, are also shown. Nonconserved bases at the CCR2 and CCR5 loci are shown in lowercase letters. Symbols are as in Figure 1. (D) PCR products validating duplications in two clones. No other mutations or rearrangements were detected in these clones. (+) is a positive control (cells treated with ZFN-224) and (−) is a negative control (cells transfected with empty plasmid). (E) Schematic of CCR2 and CCR5 loci in the wild-type and duplication alleles and Southern blot analysis. Symbols are as in Figure 1.











