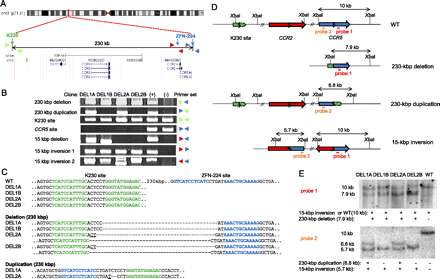
Various chromosomal rearrangements induced by ZFNs. (A) Schematic of ZFN target sites and chemokine receptor genes on the human chromosome 3 (http://genome.ucsc.edu). Note that ZFN-224 has two target sites, one at the CCR5 locus and the other at the CCR2 locus. Colored triangles indicate approximate positions of PCR primers. (B) PCR products validating various chromosomal rearrangements. (+) indicates a positive control (cells transfected with plasmids encoding two ZFN pairs) and (−) indicates a negative control (cells transfected with empty plasmid). (C) DNA sequences of PCR products. Each ZFN target site is shown in boldface letters. K230 and CCR5 target sites are shown in green and blue letters, respectively. Note the absence of the intact CCR5 (ZFN-224) site in these clones. Microhomologies are underlined and inserted bases are shown in italics. Dashes indicate deleted bases. (D) Schematic of ZFN target sites in wild-type cells and in various clones with chromosomal rearrangements. Two probes (red and orange bars) and the site recognized by the restriction enzyme, XbaI, used for Southern blot analysis are indicated. (E) Southern blot analysis of chromosomal rearrangements. Genomic DNA digested with XbaI and hybridized with probe 1 (upper) or with probe 2 (lower). A direct evidence of the 15-kbp inversion was provided by DNA sequencing (Supplemental Fig. 1).











