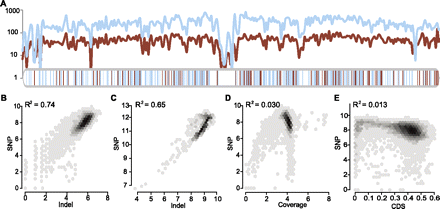
The correlation of SNP and small indel densities. (A) Parallel change of SNP and small indel density on Chr1. The density was defined to be the number of SNPs/indels per 100 Kb. (Blue curve) SNP density; (red curve) small indels. Blue and red vertical bars below show the location of large deletions and insertions, respectively. (B,C) Linear regression of SNP and indel densities in 100-Kb (B) or 1-Mb (C) sliding windows. (D,E) Linear regression of SNP density with read coverage (D) or CDS fractions (E) in a 100-Kb sliding window. All values are log2 transformed before applying regression. A near zero R2 value suggests that read coverage or CDS fractions do not contribute much to the correlation.











