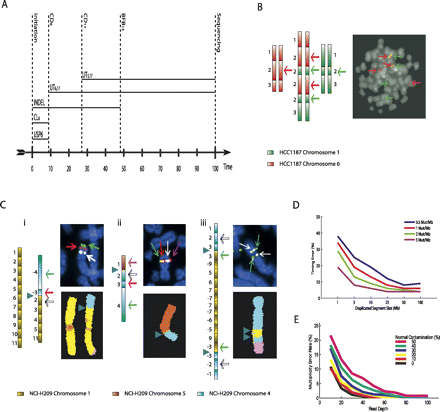
Validation. (A) The estimated timeline of rearrangement and selection events through oncogenesis relative to a molecular clock (along the horizontal axis) for the two clusters of PD3904. Events that can be timed are represented by vertical lines. Events that can only be ordered relative to these times are indicated by horizontal lines. (B,C) The predicted sequences of segments and FISH images for the two clusters of Figure 3iii,iv, respectively. Each segment is represented as a rectangle, with light to dark shading indicating the left and right ends of each segment. The number labels for each segment are as described in Figure 3 and the Supplemental Material. Green and red segments correspond to chromosomes 1 and 6 of HCC1187. Yellow, blue, and brown segments represent chromosomes 1, 4, and 5 of NCI-H209. Arrows indicate positions and luminescence of probes designed to test predicted adjacency of segments. For HCC1187, the green and red probes hybridize to segment 2 of chromosomes 1 (denoted 21) and 26, respectively. For Ci, white, red, and green probes hybridized to 21, 34, and 44, respectively. Magenta, white, red, and green probes in Cii hybridized to 15, 24, 34, and 44, respectively. Green and white probes in Ciii hybridized to 31 and 24, respectively. Breakpoints between chromosomes are represented by triangles. (D) The mean error of predicted rearrangement times. Mutations were generated at background prevalence of 0.5, 1, 2, and 5 mutations per megabase. Tandem duplications were constructed of lengths 1, 5, 10, 25, 50, and 100 Mb at random times. The mean errors of the prediction time of rearrangements from 1000 simulations are indicated. (E) The predicted error of multiplicity for normal contamination ranging from 0% to 50%, and read depth up to 100.











