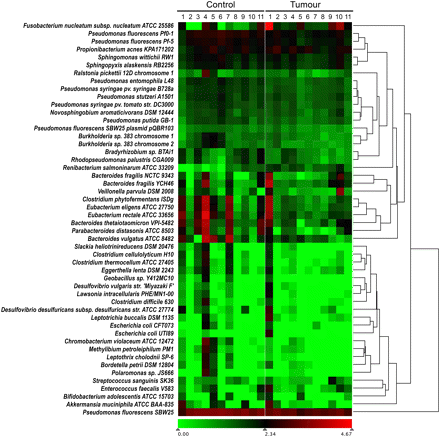
Relative abundance of microbial genomes in tumor and control specimens. Numbers of read-pairs that matched known microbial sequences were normalized according to sequencing depth for both tumor and matched normal samples. The abundance of normalized bacterial read-pairs ranged from zero to a maximum of 66,896 represented by a transition from green to red on a log10 scale. F. nucleatum sequences were present in the tumor samples at levels twofold or greater than in normal samples in nine out of the 11 subjects. The mean over abundance across all subjects was 79-fold.











