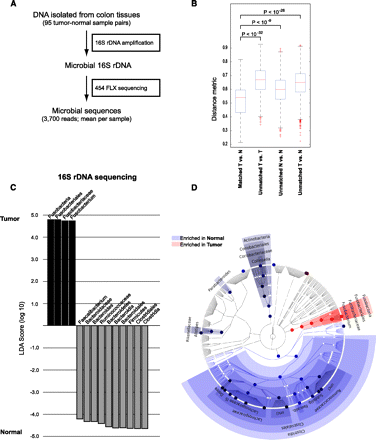
16S rDNA sequencing analysis of the colorectal cancer microbiome. (A) Schematic of experimental and computational 16S rDNA sequencing analysis workflow. (B) Beta-diversity distances calculated using phylotype relative abundance measurements between all pairs of samples demonstrate that the microbial composition of tumor/normal pairs within individuals is more highly correlated than tumor/tumor pairs, normal/normal pairs, or tumor/normal pairs from different individuals. (C) Linear discriminant analysis (LDA) coupled with effect size measurements identifies Fusobacterium as the most differentially abundant taxon in colon tumor versus normal specimens by 16S rDNA sequencing in 95 individuals. Tumor-enriched taxa are indicated with a positive LDA score (black), and taxa enriched in normal tissue have a negative score (gray). Only taxa meeting an LDA significant threshold of 4.2 are shown. (D) A cladogram representation of data in C. (Red) Tumor-enriched taxa; (blue) taxa enriched in normal tissue. The brightness of each dot is proportional to its effect size.











