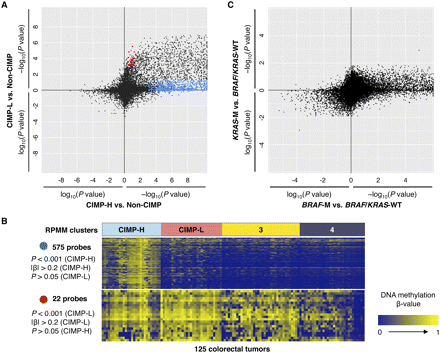
DNA methylation characteristics associated with CIMP-H, CIMP-L, BRAF-, and KRAS mutant colorectal tumors. (A) Comparison of CIMP-H- and CIMP-L-associated DNA methylation profiles. Each data point represents the log10-transformed FDR-adjusted P-value comparing DNA methylation in CIMP-H (n = 28) versus non-CIMP tumors (n = 68) (x-axis) and in CIMP-L (n = 29) versus non-CIMP tumors (n = 68) (y-axis) for each Infinium DNA methylation probe. For the probes with higher mean DNA methylation in CIMP-H or CIMP-L tumors compared to non-CIMP tumors, −1 is multiplied by log10(FDR-adjusted P-value), providing positive values. The blue and red points highlight probes that are significantly hypermethylated in CIMP-H and CIMP-L tumors compared to non-CIMP tumors, respectively. (B) Heatmap representing Infinium DNA methylation β-values for 575 CpG sites that are significantly hypermethylated in CIMP-H compared with non-CIMP tumors (top) and 22 CpG sites that are significantly hypermethylated in CIMP-L compared with non-CIMP tumors (bottom). The four DNA methylation-based subgroups are indicated above the heatmaps. A color gradient from dark blue to yellow was used to represent the low and high DNA methylation β-values, respectively. (C) Comparison of BRAF mutant- and KRAS mutant-associated DNA hypermethylation signatures in CRC. The log10-transformed FDR-adjusted P-value for each probe is plotted for tumors harboring KRAS mutations (KRAS-M) (n = 34) versus BRAF/KRAS wild-type (n = 74) (y-axis) and those containing BRAF mutations (BRAF-M) (n = 17) versus BRAF/KRAS wild-type (n = 74) (x-axis). For the probes with higher mean DNA methylation β-values in BRAF or KRAS mutant tumors compared to wild-type tumors, −1 is multiplied by log10(FDR-adjusted P-value), providing positive values.











