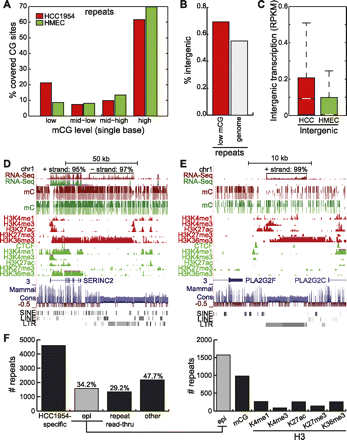
Epigenetic contribution of increased repeat expression. (A) Distribution of %mCG of each repeat-associated cytosine residue spanned by at least 10 methylC-seq reads. (B) Percentage of all repeats (gray) and of lowly methylated repeats in HCC1954 (red) that are intergenic. (C) Distribution of intergenic repeat expression in HCC1954 and HMEC, expressed in RPKM. (White line) median, (box) 25th to 75th percentiles. (D) Snapshot of repeat expression and read-through with epigenetic signatures near the SERINC2 gene. RNA-seq strand distribution shown on top. (E) Snapshot of repeat expression and read-through with epigenetic signatures near the PLA2G2F gene. RNA-seq strand distribution shown on top. (F) Number of highly expressed repeats specific to HCC1954 and the number that can be explained by at least a 50% change in epigenetic modifications or read-through transcription from nearby repeats. (Epi) epigenetic.











