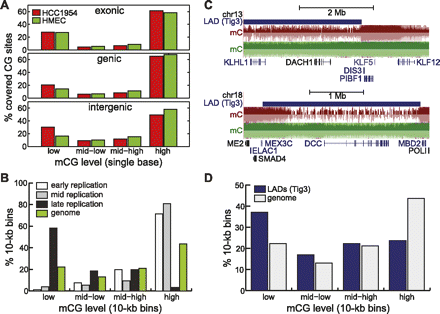
A passive model of hypomethylation. (A) Distribution of HCC1954 %mCG for exonic (top), genic (middle), and intergenic (bottom) cytosine residues. (B) Distribution of HCC1954 %mCG for 10-kb regions that are consistently early-, middle-, and late-replicating in four cell types (Hansen et al. 2009), compared to the background genome (green). (C) Snapshots illustrating the overlap of hypomethylated regions with lamina-associated domains previously mapped in Tig3 cells (Guelen et al. 2008), at the DACH1 (top) and DCC (bottom) genes. (D) Distribution of HCC1954 %mCG for 10-kb regions that are found in Tig3 lamina-associated domains (blue), compared to the background genome (gray).











