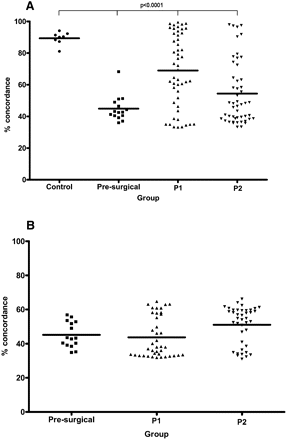
Plasma of breast cancer patients shows low SNP concordance with paired normal DNA. (A) Percentage of concordant SNP genotype calls for paired plasma and normal leukocyte DNA samples of patients and healthy controls. Percentage of concordance was significantly lower than controls in breast cancer patients (P < 0·0001, one-way ANOVA). (B) Percentage of concordant SNP genotype calls for paired plasma and microdissected tumor (available for all presurgical patients and 40 patients on follow-up; mean 47.00%; range, 31.04%–66.20%; 95% CI, 0.07–2.28). In A, concordance was lowest for the 15 preoperative primary breast cancer patients (mean, 44.88%; range, 36.00%–68.27%; 95% CI, 0.13–4.02) but remained low for the 50 patients on follow-up using both P1 (mean, 69.10%; range, 33.17%–99.44%; 95% CI, 0.21–6.51) and P2 plasma samples (mean, 54.22%; range, 33.31%–97.96%; 95% CI, 0.18–5.65). Control indicates healthy female controls; presurgical, plasma of presurgical breast cancer patients; and P1 and P2, first and second plasma samples of patients on follow-up.











