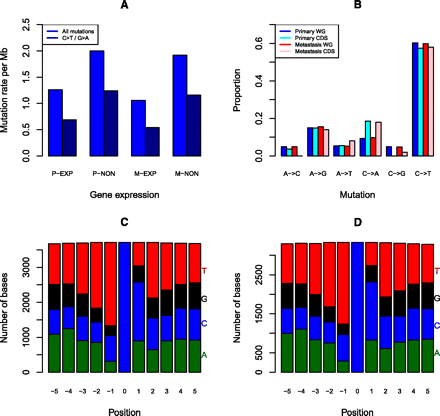
Somatic SNVs analysis in acral melanoma samples. (A) Somatic SNV mutation rates of all SNVs and C>T/G>A transitions for expressed genes in the primary (P-EXP) and metastatic tumors (M-EXP) and nonexpressed genes (P-NON and M-NON). (B) Proportion of somatic SNVs by class in the primary and metastatic tumors for the whole genome (WG) and coding regions (CDS). (C) Frequency of bases ±5 bp of whole genome C>T/G>A transitions in the primary tumors. (D) Frequency of bases ±5 bp of whole genome C>T/G>A transitions in the metastatic tumors.











