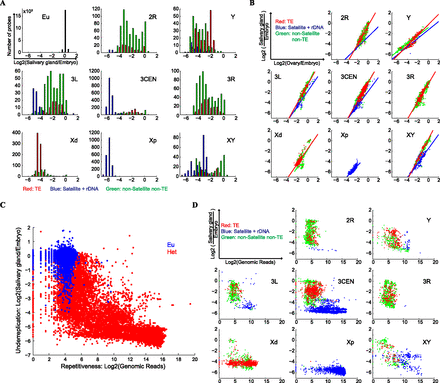
Repetitiveness of the H-probes and their underreplication in polytene chromosomes. (A) CGH analysis of polytene DNA purified from larval salivary glands and DNA from blastoderm stage embryos collected prior to tissue differentiation and chromosome polytenization. The data were normalized such that the mean log2-ratios of polytene DNA and embryonic DNA were zero for single-copy euchromatic genes (Eu). The H-probes were strongly underrepresented in the polytene DNA, and the level of underreplication varied for H-probes mapped to different chromosomal regions. (B) Comparing the degree of underreplication in salivary glands and ovaries. The colored lines are the linear fits to the data for the indicated category of H-probes. Note that the slope of linear regression fit for the satellite-like H-probes is less steep compared with the non-satellite-like H-probes. (C) Comparing sequence repetitiveness with level of underreplication in polytene chromosomes. To measure repetitiveness, sequence reads from an independent D. melanogaster genomic library were aligned to the probe sequences and the number of reads per probe was taken as an indication for repetitiveness. (Eu) Annotated euchromatic genes. (Het) H-probes. (D) H-probes from different sequence element families within individual heterochromatic regions show characteristic patterns of repetitiveness and underreplication in salivary gland polytene chromosomes.











