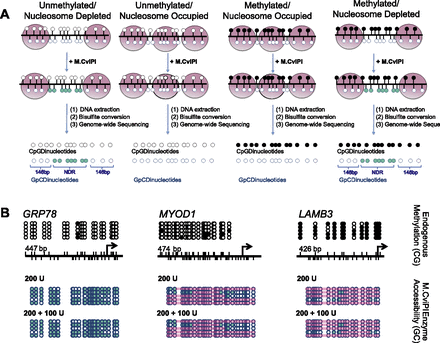
NOMe-seq can footprint a variety of chromatin structures. (A) After IMR90 cell nuclei are treated with M.CviPI, DNA is extracted, bisulfite-converted, and sequencing is performed. DNA methylation status is obtained from CpG dinucleotides, and nucleosome occupancy information is gained from the inaccessibility of the M.CviPI methyltransferase to GpC dinucleotides. The combination of DNA methylation and nucleosome occupancy data can reveal four distinct chromatin signatures: unmethylated and nucleosome-depleted, unmethylated and nucleosome-occupied, methylated and nucleosome-occupied, and methylated and nucleosome-depleted. (Black circles) Methylated CpG sites; (teal circles) accessible (methylated) GpC sites. (B) We found that 200 units of M.CviPI for 7.5 min followed by a boost of 100 units accurately revealed an NDR upstream of the TSS of HSPA5 (also known as GRP78), an active CGI promoter, while also showing that the polycomb repressed MYOD1 CGI promoter and methylation-silenced CpG-poor LAMB3 promoter were occupied by nucleosomes and inaccessible to M.CviPI, as expected. M.CviPI-inaccessible regions greater than 146 bp are covered by a pink rectangle indicating nucleosome occupancy. PCR amplicon sizes: HSPA5–447 bp, MYOD–474 bp, and LAMB3–426 bp.











