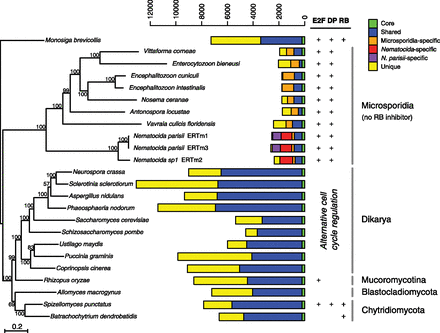
Phylogeny and ortholog content of Microsporidia and Fungi. The phylogeny was inferred by maximum likelihood using RAxML (Stamatakis 2006) based on the concatenated amino acid sequences of 53 single-copy orthologs shared by all taxa. Bootstrap values above 50% are listed above respective nodes. Ortholog counts are shown as bar graphs to the right of the tree and are divided into six categories: core (green, found in all genomes), shared (blue, found in at least two genomes excluding other Microsproridia and Nematocida categories that follow), Microsporidia-specific (orange), Nematocida-specific (red), N. parisii–specific (purple), and unique (yellow). The presence of E2F, DP, and RB orthologs is indicated in table on the right, with E2F and DP activators in green, and the RB inhibitor in red. Dikarya cell cycle regulation circuitry is evolutionarily distinct from the RB–E2F pathway (Cao et al. 2010).











