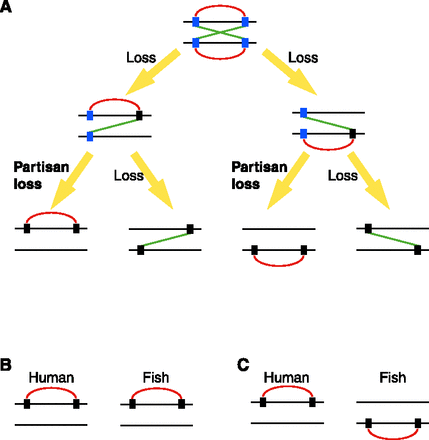
PPIs in orthologous/paralogous paralogons in vertebrate genomes. Vertical and horizontal lines represent genes and chromosomes, respectively. Blue and black rectangles show extant-paired ohnologs and singletons, respectively. Red and green lines indicate cis- and trans-PPIs, respectively. (A) Gene loss patterns of cis-linked genes after WGD. Partisan loss can be observed when the second gene loss occurs in a paralogon where the first gene loss occurred. (B) When we observe conserved cis-linked genes in an orthologous paralogon between human and fish, it is difficult to distinguish an independent partisan loss from a cis-PPI derived from a common ancestor. (C) When we observe conserved cis-linked genes in a paralogous paralogon between human and fish, this pattern is strong evidence of independent partisan loss after speciation between human and fish.











