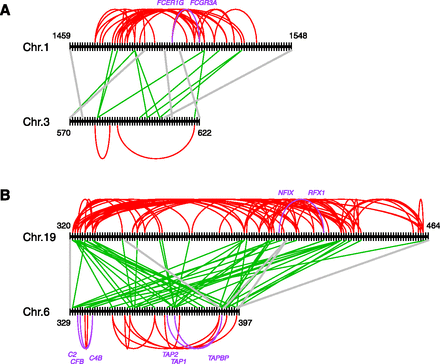
Cis- and trans-PPIs between genes in human paralogons. Black vertical and horizontal lines represent genes and chromosomes, respectively. The numbers beside chromosomes indicate gene ordinal numbers along the chromosome. (Bold gray lines) Homology relationships between extant-ohnolog pairs. Red and green lines indicate cis- and trans-PPIs between genes in a human paralogon, respectively. Purple lines and gene names denote cis-PPIs previously identified as interacting gene clusters related to “immune response” (Makino and McLysaght 2008). In the case of physical links in paralogons, it is possible to identify cis-PPIs over a wider range compared with searches within a fixed base-pair distance (Makino and McLysaght 2008). There are many more cis-PPIs compared with trans-PPIs in (A) paralogon ID 24 (27 cis and 10 trans) and in (B) paralogon ID 507 (69 cis and 46 trans) (Supplemental File 1). Even after collapsing tandem duplicated genes, the PPIs of singletons are enriched in cis (ID: 24: 18 cis and 8 trans; ID 507: 60 cis and 34 trans).











