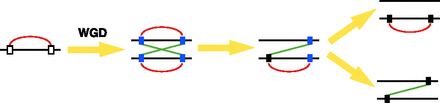
Gene losses after whole genome duplication (WGD). Rectangles and horizontal lines represent genes and chromosomes, respectively. Red and green lines indicate cis- and trans-protein-protein interactions (PPIs) between proteins encoded by singletons or extant-paired ohnologs, respectively. (White rectangles) Genes prior to WGD. Blue and black rectangles show extant-paired ohnologs and singletons, respectively. Following WGD, all interacting gene clusters will be perfectly duplicated, resulting in exactly equal numbers of cis- and trans-PPIs. The first gene loss can occur at any locus. Gene loss that reverts the second gene to single copy will result in either retention of a cis- or a trans-PPI. If gene loss is neutral, then both scenarios should occur with equal frequency.











