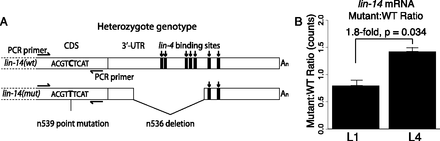
lin-14 mRNA abundance in animals heterozygous for a 3′ UTR deletion. (A) Schematic of wild-type and mutant lin-14 alleles in the heteroallelic configuration. The n536 deletion is confined to the 3′ UTR and eliminates five lin-4 binding sites. The n539 point mutation is used to distinguish mutant from wild-type transcript; reverse transcription and PCR primers were selected using regions around the point mutation that are identical in both alleles, generating products of identical length and differing only at a single internal position. (B) Abundance ratios for wild-type and mutant alleles in L1 and L4 larvae, as measured by Illumina sequencing. Individual ratios were measured as the simple ratio of mutant to wild-type reads, error bars represent standard error of the mean, P-value is from a two-sided t-test. Equivalent data derived using Sanger sequencing is shown in Supplemental Figure S3.











