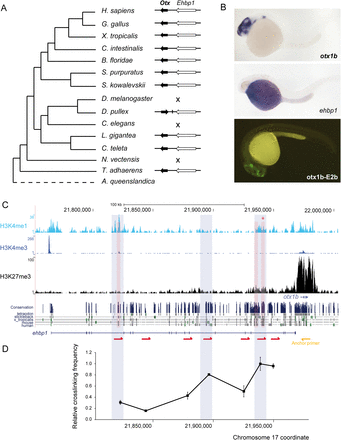
Functional characterization of the GRB of otx1b-ehbp1 in zebrafish. (A) Phylogenetic distribution of the GRB across the studied metazoan species. The GRB was not conserved in D. melanogaster, C. elegans, and N. vectensis. (B) Top and middle panels are zebrafish embryos at 24 hpf showing the expression of otx1b and ehbp1 genes. otx1b is detected in the anterior brain, while ehbp1 is expressed in the yolk. (Lower panel) GFP expression promoted by an enhancer located within the intron of ehbp1 (asterisk in C). This enhancer is active in most tissues expressing otx1b. (C) Distribution of H3K4me3, H3K4me1, and H3K27me3 tracks along the otx1b-ehbp1 GRB in 24-hpf zebrafish embryos. H3K4me1 peaks tested for enhancer activity in zebrafish stable transgenic lines are shaded in red. The regions that physically interact in 3C assays with the otx1b promoter are shaded in blue. H3K27me3 distribution indicates that otx1b, but not ehbp1, has tissue-specific expression; hence, H3K4me1 enhancers located in ehbp1 introns are likely acting on otx1b. (Below) conservation track from the UCSC Genome Browser. (D) Graph showing a 3C experiment to detect interaction between different ehbp1 intronic regions and the otx1b promoter in 24-hpf embryos. A fixed primer (yellow arrow) was set at the otx1b promoter, and seven regions were assayed for interaction with that promoter using different primers (red arrows) distributed along the ehbp1 intronic genomic area. The highest cross-linking frequency value is set to 1. Error bars indicate standard error (n = 3).











