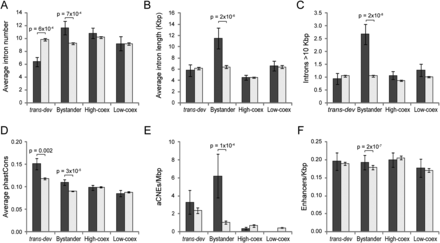
Different functional and evolutionary features of intragenic sequences of conserved pairs. (A) Average intron number for each gene type. (Dark gray) Conserved pairs; (light gray) control set. Putative human bystander genes (nondevelopmental genes associated with trans-dev genes in conserved pairs) have significantly more introns on average than the control set (P = 7 × 10−4, KS test), whereas associated trans-dev genes have fewer (P = 6 × 10−4, KS test). Genes in highly coexpressed or lowly coexpressed conserved pairs have similar average intron numbers than the control set. (B) Average intron length (in Kbp). Putative bystander genes have significantly longer introns than the other genes (P = 2 × 10−6, KS test). (C) Average number of introns longer than 10 Kbp. Bystander genes have nearly three times more long introns than other genes (P = 2 × 10−8). (D) Average mammalian-wide phastCons score of intronic regions of different types of conserved gene pairs. Both trans-dev and bystander genes from conserved pairs show higher intronic sequence conservation than the nonconserved genes (P = 0.002 and P = 3 × 10−5, KS test), whereas conserved highly coexpressed nondevelopmental gene pairs show higher conservation than the control set at the intergenic region. (E) Density of ancient conserved noncoding elements (aCNEs) per Mbp in introns of different types of conserved gene pairs. (F) Density of functionally defined strong enhancers per Kbp in introns of different types of conserved gene pairs. Error bars correspond to standard errors.











