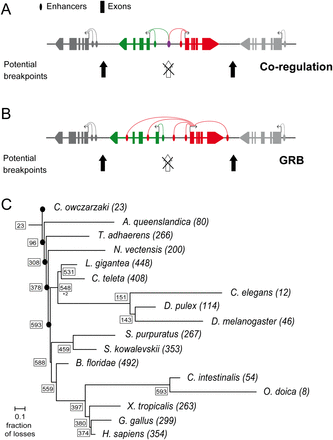
Functional causes of conserved microsynteny and phylogenetic distribution of conserved gene pairs. (A) (Coregulation) Two neighbor genes share one or more regulatory elements. Chromosomal breakpoints in the intergenic region of the two genes are opposed by selection since it would affect the coordinated expression of the two genes. (B) Genomic regulatory block (GRB): A bystander gene (green) contains regulatory elements in its introns that target a neighbor gene (red), often a trans-dev gene with key regulatory roles in animal development. The breakage of the association would result in affected regulation of the trans-dev gene. (Based on Becker and Lenhard [2007]). (C) Consensus phylogeny of the studied metazoans, showing the total number of CAMPs in each species (in parentheses), and the minimum number of pairs at each node inferred by parsimony (in boxes). Branch lengths correspond to the fraction of gene pairs lost out of the total pairs present in the last common ancestor. Black dots at basal nodes indicate that branch length could not be estimated.











